From feathered dinosaurs to woolly mammoths, countless animals that no longer live have come to pervade modern popular culture. Snuck into a world of dragons, witches, and ice zombies in Game of Thrones (or A Song of Ice & Fire), the dire wolf recently experienced a slightly confusing reintroduction into the beloved pantheon of extinct megafauna, as it may appear to some to be mythical by association. Dire wolves were very much real, and we have known about them since the mid-nineteenth century. These charismatic canids roamed the Americas and parts of eastern Asia for more than 100,000 years, going extinct early into the Holocene around 9,000 to 10,000 years ago (BP).What were they really like, and why did they go extinct? In answering these questions, we can also begin to explore the diverse ways in which information is gleaned from fossils.
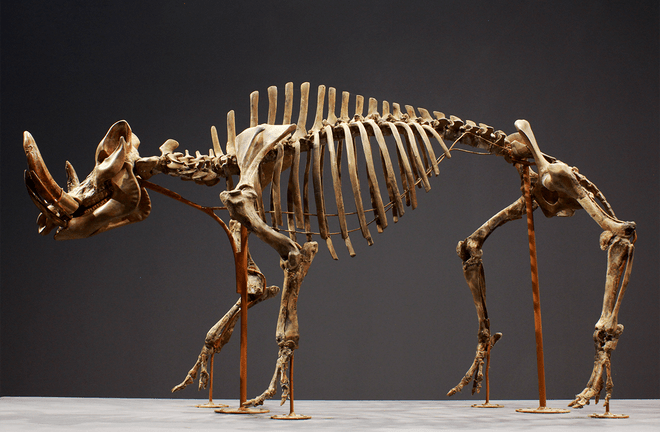
We start our story in the La Brea Tar Pits in Los Angeles, USA, the largest discovered fossil site containing dire wolves, numbering beyond 4,000 individuals. Tar pits form from crude oil seeping up to the earth surface and congealing into asphalt. The La Brea Tar Pits contained the remains of thousands of animals representing several extant and extinct species that fell into and became deposited in the asphalt between 40,000 and 8,000 BP. Excavations at La Brea began in 1913, and the asphalt-preserved fossils continue to be used to reimagine not just the organisms themselves, but also the world that they inhabited.

A predator and its prey
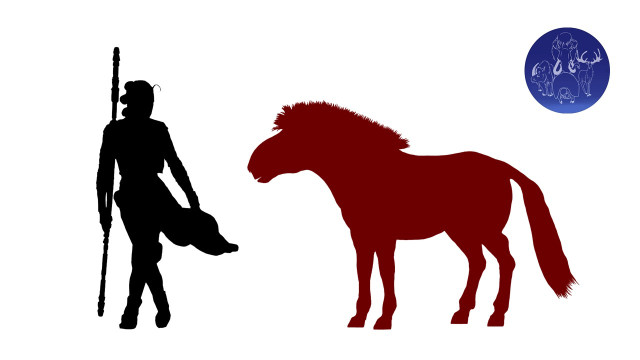
One way to get to know an apex predator is through its preferred prey. Many chemical elements exist as multiple, stable isotopes that carry different numbers of protons in their nuclei. For instance, carbon exists as the common type 12C and the much rarer 13C, while nitrogen exists as the common and rare 14N and 15N, respectively. The stable isotope ratios 13C/12C and 15N/14N vary across different kinds of plants based on their environment and how they incorporate the elements from the atmosphere, and in animals depending largely on their diets. Stable isotope compositions of biological tissues such as bone can reconstruct ancient diets and food webs. A 2007 study first identified how isotope ratios in modern wolves are related to that in their prey, beavers and moose. Applying these findings to isotope ratios in the bones of dire wolves and several candidate herbivore prey species from the La Brea Tar Pits, the authors inferred that dire wolves primarily preyed on horses, and to a lesser extent on mastodon, sloth, and grazers such as bison.

We know dire wolves were likely pursuit predators who chased down their prey and attacked with their jaws alone, based on our knowledge of extant wolves and other canids. This has also been verified through fossil evidence. The authors of a 2017 study predicted and found that patterns of skeletal injuries differed between La Brea dire wolves and sabretooth cats, which were likely ambush predators. Sabretooth cats had frequent spinal injuries from grappling with their prey, while dire wolves tended to have high rates of trauma across all four limbs, consistent with chasing down prey.
Clues to extinction
Patterns and frequencies of tooth wear and damage have been found in living animals to be informative of feeding behaviors. A 1993 study brought this understanding to the study of several La Brea carnivores, including dire wolves. The authors found a high frequency of tooth breakage across the carnivores, likely from tooth-bone contact, suggesting greater utilization of carcasses compared to modern carnivores. Was this due to a decline in available herbivore prey in the late Pleistocene, eventually leading to the extinction of these predator species?
To answer this question, a 2002 study compared dire wolves preserved in two La Brea pits dated to ~15,000-14,000 BP and ~13,000-12,000 BP, respectively, and found that carcass utilization actually decreased over this time span. Times appear to have been especially tough around ~15,000-14,000 BP, leading to more complete consumption of prey to avoid scavenging by other competing predators. The subsequent decline in this behavior may reflect dwindling predator densities as dire wolves and other carnivores approached extinction. A 2015 study found a similar temporal pattern, among other findings.
The study investigated several measures of morphological variation using the remains of 83 dire wolves as well as 97 sabertoothed cats from La Brea, spanning the period between 40,000 and 12,000 BP. One of these measures, fluctuating asymmetry, quantifies (to a reasonable extent) developmental instability, the tendency of traits to deviate from the target phenotype. Symmetrical traits such as the left and right sides of the face are controlled by the same genetic and developmental pathways, and any asymmetry implies developmental instability, typically caused by some form of external stress. The study found increasing fluctuating asymmetry in dire wolves over time from 40,000 BP until ~15,000-14,000 BP, possibly in response to environmental stress associated with rapid climatic transitions during that period. This was followed by a decline in fluctuating asymmetry 13,000-12,000 years ago, mirroring the changes in tooth wear and breakage over the same interval. Together, these studies imply that climatic instability leading up to the Holocene put dire wolf populations under stress, likely through disrupting populations of their herbivore prey, which they never completely recovered from. Humans are also thought, quite contentiously, to have played some role in Pleistocene megafaunal extinctions.
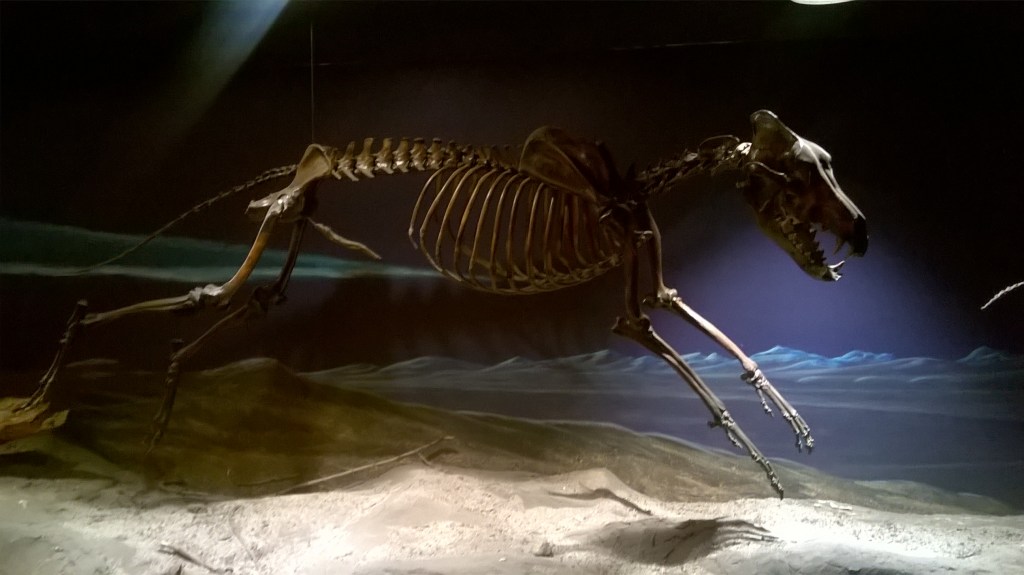

A wolf by any other name
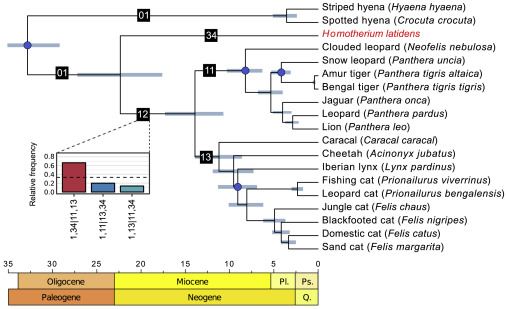
We end with a reflection on the evolutionary relationships of the dire wolf. Over the last two decades, enormous advances have been made in techniques to extract preserved DNA from ancient remains. Ancient DNA provides snapshots of the evolutionary past, and in the case of extinct animals, a way to infer their relationships with extant taxa. A 2021 study looked at ancient DNA from dire wolf remains, and found that dire wolves were in fact distantly related to extant gray wolves. The two species had diverged from a common ancestor more than 5 million years ago, and look so similar largely because they convergently adapted to lives of pursuit predation. To place this in context, jackals, coyotes, and gray wolves are more closely related to each other than to dire wolves.
While modern canids like grey wolves and coyotes are known to sometimes interbreed, the study found that this was not the case between dire wolves and grey wolves. Hybridization leads to the introduction of new genetic diversity, which can often help a species adapt to changing conditions. Dire wolves were not beneficiaries of this process as they disappeared along with the prey species they had been adapted to hunting and eating.
Humans have likely been coming across and pondering fossilized remains of extinct animals for thousands of years. Records of such encounters are thought to persist today in the form of mythological beasts reimagined from the fossils, such as giants, griffins, and dragons. Modern day reconstructions from fossil remains are, as we have just seen, much less fanciful, but far more exciting in what they tell us about ecosystems and episodes in evolutionary history that have been lost to time.
Written by Ornob Alam (Twitter @genomeinquirer)
Edited by Jan Freedman (Twitter @JanFreedman)
Ornob is a graduate student in Michael Purugganan’s lab at New York University. His dissertation research examines the demographic and evolutionary history of domesticated Asian rice in the context of past climate change and human migrations.
Further reading:
Binder, W. J., Thompson, E. N., and Valkenburgh, B.V. 2002. “Temporal variation in tooth fracture among
Rancho La Brea dire wolves,” Journal of Vertebrate Paleontology, vol. 22, no. 2, pp. 423–428, Jul. 2002, doi: 10.1671/0272-4634(2002)022[0423:TVITFA]2.0.CO;2. [Abstract only]
Broughton, J. M., and Weitzel, E. M, 2018. “Population reconstructions for humans and megafauna
Suggest mixed causes for North American Pleistocene extinctions,” Nature Communications, vol. 9, no.
1, Art. no. 1, Dec. 2018, doi: 10.1038/s41467-018-07897-1. [Abstract only]
Brown, C., Balisi, M., Shaw, C. A., and Van Valkenburgh, B. 2017. “Skeletal trauma reflects hunting
behaviour in extinct sabre-tooth cats and dire wolves,” Nature Ecology & Evolution, vol. 1, no. 5, Art. no.
5, Apr. 2017, doi: 10.1038/s41559-017-0131. [Abstract only]
Fox-Dobbs,K. F. D., Bump, J. K. B. K., Peterson, R. O. P. D., Fox D. L. F., and P. L. K. L. Koch, P. L. K. 2007.
“Carnivore-specific stable isotope variables and variation in the foraging ecology of modern and ancient
wolf populations: case studies from Isle Royale, Minnesota, and La Brea,” Canadian Journal of Zoology,
doi: 10.1139/Z07-018. [Abstract only]
Goswami A., Binder. W. J., Meachen, J., and O’Keefe, F. R. 2015. “The fossil record of phenotypic
Integration and modularity: A deep-time perspective on developmental and evolutionary dynamics,”
Proc Natl Acad Sci U S A, vol. 112, no. 16, pp. 4891–4896, Apr. 2015, doi: 10.1073/pnas.1403667112.
Perri, A. R., et al., 2021. “Dire wolves were the last of an ancient New World canid lineage,” Nature, vol.
591,No. 7848, Art. no. 7848, Mar. 2021, doi: 10.1038/s41586-020-03082-x. [Abstract only]
Stewart, M., Carleton, W. C., and Groucutt, H. S. 2021. “Climate change, not human population growth,
correlates with Late Quaternary megafauna declines in North America,” Nature Communications, vol.
12, no. 1, Art. no. 1, Feb. 2021, doi: 10.1038/s41467-021-21201-8. [Abstract only]
Vanvalkenburgh, B., and Hertel, F. “Tough times at la brea: tooth breakage in large carnivores of the late
pleistocene,” Science, vol. 261, no. 5120, pp. 456–459, Jul. 1993, doi: 10.1126/science.261.5120.456.