When I first started doing ancient DNA way back in two thousand and [polite cough], every base pair was a win. Nature papers were published with less sequence than you’d now expect a fresher to produce in their first lab practical. The exciting thing about being in a new and expanding field was the way in which new records were constantly being broken, week by week. The first ancient mitochondrial genome. The first middle Pleistocene DNA, the first nuclear DNA, the first time-stamped dataset, the first Neanderthal genome. All of these ground breaking studies were published while I was a young PhD and post-doc. All of them received enormous amounts of publicity, and rightly so. Each pushed the boundaries in a way that only new research techniques can. My very first paper was published in a reasonably high profile journal and consisted of only a couple of kilobases of data. You’d be lucky to get that published in a Christmas round robin these days.
I distinctly remember lab meetings as a young PhD where serious discussions happened about whether we would ever be able to produce an ancient nuclear genome. And this was amongst other ancient DNA specialists. We just didn’t know. My own early work painstakingly teasing a few base pairs of ancient mitochondrial DNA out of well-preserved sabretooth fossils, while momentous at the time, definitely got my mind spinning over whether one day we would ever get a complete nuclear genome from any extinct cat. While the mitochondrial DNA can tell us about maternal lineages and some demography, the nuclear DNA is the real deal. This is the DNA that is inherited from both mother and father and codes for practically everything that makes up an organism. Whether hair is long or short, patterned or plain, dark or light. How each part of the organism integrates with the whole. The nuclear DNA is the blueprint for life itself and can tell us about phenotype but also about population change, population size, and evolutionary selection pressures, amongst many other things.
Periodically, as sequencing technologies have progressed, I’ve revisited the idea of getting complete genomes from ancient cats. Earlier this year we published the first ancient genomes from a range of extinct lions [de Manuel, Barnett et al.]. But throughout all this, one species has exerted a siren call over me. My favourite extinct species, a ghost of the ice age steppe, and shoe-in for most impressive cat ever to have lived. I’m talking of course about our old friend the Scimitar cat, Homotherium latidens.
We have recently [Westbury, Barnett et al. 2020] published the first ancient nuclear genome of a sabretooth cat. The work is intimately tied to the work I did on ancient lion genomes while working as a postdoc in Copenhagen on a Marie Curie research fellowship. The MCRF work had me focusing on lion genomics, including that of the cave lion (Panthera spelaea). To get good samples, I relied on my friend Dr Grant Zazula who is the state palaeontologist for the Yukon government. He was able to send me some fresh cave lion bones straight out of the Yukon permafrost. These ancient bones had been on ice for 20 thousand years and had excellent DNA preservation. Getting genomes from ancient bones involved first grinding up small bits of them and applying various bucket chemistry tricks to remove the protein/fat/mineral content to leave the DNA behind. This DNA then gets chemically joined to artificially produced DNA that incorporates various binding sites and a unique DNA barcode that allows each fragment to be reliably separated out. This soup of barcoded DNA bits is known as a library, and can be amplified by standard PCR methods and then run on a pyrosequencing machine to produce billions of reads of DNA that represent all the DNA in your extract. Once the initial library is created it will be sequenced as a matter of course to check what percentage of the extracted DNA is actually from the organism you are interested in versus background DNA from contaminants/humans/bacteria/fungi. Obviously the higher percentage of organismal DNA you have to start with the easier (and more importantly, cheaper) it will be to generate enough sequence data to stitch together a complete genome. For some of the cave lion bones Grant sent me, we got phenomenal levels of endogenous content, and those samples ended up on the lion genome paper. One of the samples was a little different though. When I got back that first initial library sequencing, I was sitting on my sofa at home, using my laptop to check how much lion there was in the mix. This involved comparing all the tiny, barcoded fragments to a reference standard of a modern lion mitochondrial genome (although we were ultimately interested in the nuclear genome, comparing how much mtDNA is in the library is easier to do and gives an idea of overall endogenous content that is reliable enough). While sitting and looking at all the snaking lengths of A, G, C, and T in their multi-coloured formatting I noticed something that made me sit bolt upright and nearly knock my laptop off my knees. One tiny corner of the mtgenome of the lion is seared into my mind nearly as deeply as the opening lines of Genesis. The sequence of lion ATP8 was part of the very first lion DNA I ever sequenced for my PhD and I long ago lost count of how many hours I have spent reading, and rereading, the unique sequence of this gene to check and double check that my results were good. It’s as known to me as my first address, my mother’s maiden name, my first pet. I’m never going to forget it. What shocked me into nearly dropping my laptop off my lap is that I recognised that the library sequence I had generated was not cave lion ATP8 but belonged to another cat that over the years I had known almost as intimately. The absence of just three basepairs at a critical point in the ATP8 sequence made all the hairs on the back of my neck stand to attention for I knew that only one other species had that particular deletion: Homotherium.
This bone was not a cave lion. It was a scimitar cat. One of only a handful ever recovered from the Yukon region. And what’s more it was phenomenally preserved. I think I made the decision there and then that we had to try and sequence this sample more fully and get the total nuclear genome out. Over the next six years, that’s exactly what we did, thanks to the generosity and enthusiasm of the Copenhagen Centre for GeoGenetics leader Tom Gilbert and many working members of the group. To name just two, Dr. Marce Sandoval-Velasco was crucial in generating more libraries of Homotherium after my fellowship had ended. Dr. Mick Westbury had the skills and enthusiasm to take the raw genomic data Marce and I generated, and piece it back together into a coherent whole. He also had the talent to apply the latest genomic analyses to the dataset and finally resolve some of the mysteries that shrouded Homotherium.
We were able to use the nuclear genome in a number of interesting ways. First of all, we were able to give an accurate date for when Homotherium, and by extension all the sabretooh Machairodontinae subfamily separated from the modern cats. Palaeontology had placed Homotherium, Smilodon, Xenosmilus and a bunch of other genera on their own twig of the cat family tree. We had previously estimated the separation using the mitochondrial genome but this can give false inferences if there has been any hybridisation (as seen extensively in modern cats, and our own human lineage) or sex-biased dispersal (e.g. if females stay in one place and males travel long distances between groups, as seen in lions). The nuclear data gave a fairly tight estimate of 22.5 million years (also within the bounds estimated from the mitochondrial genome previously) which puts the split pretty squarely at the Oligocene/Miocene boundary or beginning of the Miocene. Immediately, this suggests that the split between machairodonts and felines could be part of larger ecological processes happening at this geological turning point.
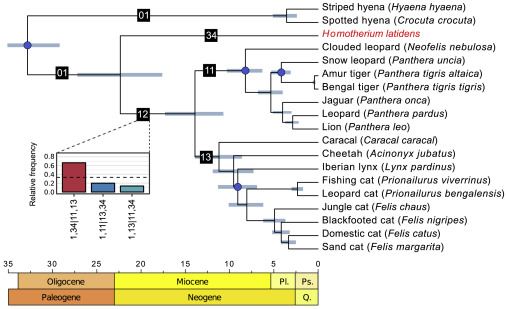
A phylogenetic tree based on nuclear data showing the position of Homotherium between the felinae and the Hyaenidae.
Tied into this, we were very interested in testing whether we could detect any signs of hybridisation between Homotherium and other felids that it would have encountered when alive. Pleistocene Homotherium ranges would have overlapped with cave lions (Panthera spelaea spelaea), American lions (Panthera spelaea atrox) leopards (Panthera pardus), tigers (Panthera tigris), and jaguars (Panthera onca), showing how global its distribution was. We see signs in the nuclear genome of modern cats that there has been admixture between lions and snow leopards, lions and tigers, lions and jaguars, and various combinations of their ancestral lineages. So, clearly big wild felids are not super discerning in who they mate with and if we could find signals of admixture between sabretooths and Panthera cats that could tell us something pretty interesting. However, in complete contrast to the Panthera cats we found no signal of any hybridisation in the Homotherium genome. It seems the scimitar cat either kept completely to itself, or any signs have been completely purged from the genome after the fact. Sad to think we never had any baby sabre-toothed lions.
Using some fancy statistical inference methods we were also able to look at demographics (population change) in Homotherium. The basic theory is that population size and genetic diversity are intimately linked (this makes intuitive sense, the larger the population the more mutations are likely to creep in). Reconstructing this from the nuclear genome of a single individual sounds strange, but remember that each genome contains information on the diversity of the mother and father as we inherit one copy of each gene from both. The appearance of heterozygosity (where DNA differences between the gene from mother and father are found) tells us about genetic diversity and hence population size. Intriguingly, despite being just about the rarest species in the Pleistocene fossil record, Homotherium seems to have had an effective population size comparable to lion, leopard, jaguar and tiger, and much greater than the cheetah or Iberian lynx. Something doesn’t add up. Maybe it’s just that because fossil cats are difficult to tell apart from most of their bones that there has been gross misidentification and all those missing Homotherium have been shut away in drawers full of cave lion or Smilodon.
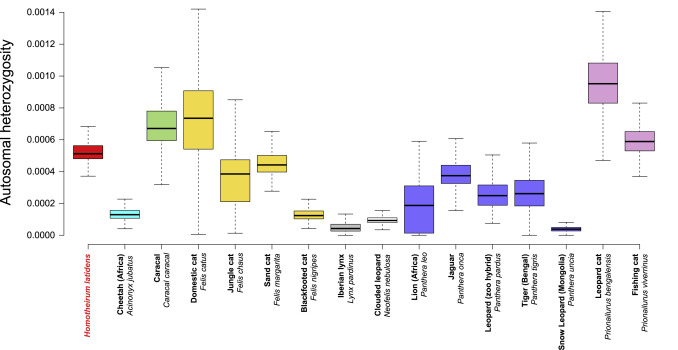
Graphic representation of nuclear autosomal heterozygosity (proxy for population size) compared between Homotherium and some species of living cat.
Another potential explanation is taphonomy. Maybe Homotherium just mostly lived in places where fossilisation was exceptional and the few scraps we have are from very lost individuals.
However, the most likely explanation is that Homotherium simply lived a very mobile life over its enormous range and this inflated the genetic diversity present in its genome.
To me, the most fascinating aspect of our genomic work has been our attempt at getting information on the phenotype from our long dead gene reads. At the moment this feels very primitive, but like all first steps it has the potential to go somewhere wonderful. We looked at which regions in the Homotherium genome were under positive selection (basically, which parts of the genome have changed and produced functional differences when compared to the same part of the genome in closely related species). When we did this we saw that genes involved with some pretty interesting things seem to have been pushed hard during the scimitar cats evolution. There were genes involved with vision and circadian rhythm, strongly suggesting that Homotherium was a diurnal hunter i.e. one that was active during the daytime and needed exceptional eyesight. There were genes involved with fat-burning, oxygen processing, respiration, and the vascular system pointing towards an active lifestyle of fast pursuit of prey similar to the cheetah or wolf, rather than the ambush hunting style of most cats. There were tantalising hints of something interesting going on in genes associated with increased sociality in model organisms, maybe pointing to Homotherium as a pack animal that could live in small hunting groups.
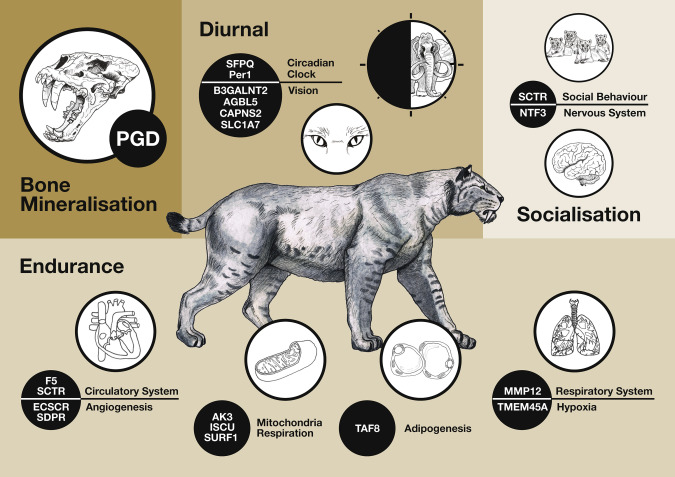
A graphical representation of which Homotherium genes we found under positive selection and what aspects of biology they are associated with in model organisms.
Unfortunately we weren’t able to recover to a decent standard some of the genes we know are involved with coat colour and appearance in cats. Genes like MC1R, ASIP, SLC45A2, TYR, FGF5 could have given us amazing insights into the life appearance of sabrecats, which would do so much to place them into their extinct niche. Modern big cats are almost entirely defined by their pelage (their skeletons are practically identical, with only minor differences) and the difference between lion and tiger is nearly all in the mane or stripes.
So, there it is. 22 year old me is faintly impressed by 40 year old me, in having a large part to do with the first sabretooth genome. May all of you stick around long enough to see some of your own dreams come to life.

A wonderful reconstruction by Velizar Simeonovski of a pair of Homotherium latidens hunting a caballine horse during the boreal winter daytime. Painting was commissioned by us to showcase some of the inferences of our paper.
Written by Ross Barnett (@DeepFriedDNA)
Further Reading:
Anton, M., M. J. Salesa, A. Turner, A. Galobart, and J. F. Pastor. 2009. “Soft tissue reconstruction of Homotherium latidens (Mammalia, Carnivora, Felidae). Implications for the possibility of representations in Palaeolothic art.” Geobios no. 42 (5):541-551. [Abstract]
Barnett, R. 2014. “An inventory of British remains of Homotherium (Mammalia, Carnivora, Felidae), with special reference to the material from Kent’s cavern.” Geobios no. 47 (1-2):19-29. [Full Text]
Barnett, R. 2019. The Missing Lynx. London: Bloomsbury.[Book]
Barnett, R., I. Barnes, M. J. Phillips, L. D. Martin, C. R. Harington, J. A. Leonard, and A. Cooper. 2005. “Evolution of the extinct sabretooths and American cheetahlike cat.” Current Biology no. 15 (15):R589-R590.[Full Text]
Barnett, Ross, Michael V. Westbury, Marcela Sandoval-Velasco, Filipe Garrett Vieira, Sungwon Jeon, Grant Zazula, Michael D. Martin, Simon Y. W. Ho, Niklas Mather, Shyam Gopalakrishnan, Jazmín Ramos-Madrigal, Marc de Manuel, M. Lisandra Zepeda-Mendoza, Agostinho Antunes, Aldo Carmona Baez, Binia De Cahsan, Greger Larson, Stephen J. O’Brien, Eduardo Eizirik, Warren E. Johnson, Klaus-Peter Koepfli, Andreas Wilting, Jörns Fickel, Love Dalén, Eline D. Lorenzen, Tomas Marques-Bonet, Anders J. Hansen, Guojie Zhang, Jong Bhak, Nobuyuki Yamaguchi, and M. Thomas P. Gilbert. 2020. “Genomic Adaptations and Evolutionary History of the Extinct Scimitar-Toothed Cat, Homotherium latidens.” Current Biology. [Full Text]
de Manuel, M., R. Barnett, M. Sandoval-Velasco, N. Yamaguchi, F. G. Vieira, M. L. Zepeda Mendoza, S. Liu, M. D. Martin, M. H. S. Sinding, S. S. T. Mak, C. Carøe, S. Liu, C. Guo, J. Zheng, G. Zazula, G. Baryshnikov, E. Eizirik, K. P. Koepfli, W. E. Johnson, A. Antunes, T. Sicheritz-Ponten, S. Gopalakrishnan, G. Larson, H. Yang, S. J. O’Brien, A. J. Hansen, G. Zhang, T. Marques-Bonet, and M. T. P. Gilbert. 2020. “The evolutionary history of extinct and living lions.” Proceedings of the National Academy of Sciences of the United States of America no. 117 (20):10927-10934.[Full Text]
Meachen, Julie A. 2017. “Ancient DNA: Saber-Toothed Cats Are the Same Beasts After All.” Current Biology no. 27 (21):R1165-R1167. doi: 10.1016/j.cub.2017.09.024.[Full Text]
Paijmans, J. L. A., R. Barnett, M. T. P. Gilbert, M. L. Zepeda Mendoza, J. W. F. Reumer, J. de Vos, G. Zazula, D. Nagel, G. Baryshnikov, J. A. Leonard, N. Rohland, M. Westbury, A. Barlow, and M. Hofreiter. 2017. “Evolutionary history of sabre-toothed cats based on ancient mitogenomics.” Current Biology no. 27 (21):3330-3336.[Full Text]
Widga, C., T. L. Fulton, L. D. Martin, and B. Shapiro. 2012. “Homotherium serum and Cervalces from the Great Lakes Region, USA: Geochronology. morphology and ancient DNA.” Boreas no. 41 (4):546-556.[Abstract]


Pingback: G(e)nomic Wisdom — TwilightBeasts – Pershspective
When you are removing the protein content do you keep any of the protein content for sequencing?
The sequencing of ancient proteins has been following the trend of that of ancient DNA but with a time lag; it is going to be more important in future because of it’s ability to last for much longer periods of time than DNA.
It doesn’t give as much information as the DNA but still it would be interest to have sequenced proteins of Homotherium to compare with fossils of even older cats.
Already it has been demonstrated that proteins of different species of Stephanorhinus differ and when people start looking at fossil elephants or fossil shark teeth then things are going to get very interesting.
Hi LeeB, yes this is a good point. Generally for ancient DNA sequencing the protein fraction gets digested with proteinase K so that it doesn’t get co-extracted and interfere with the downstream applications. Copenhagen has one of the best ancient protein groups working alongside/within GeoGenetics https://globe.ku.dk/research/evogenomics/cappellini-group/
https://globe.ku.dk/research/evogenomics/collins-group/
They did the Stephanorhinus research you allude to.
Theoretically, it would be fairly simple to keep some of the protein elution from extraction for protein analysis, but this wasn’t done for our Homotherium work because no one was doing a protein study on ancient cats at that time. There is also the potential for non-destructive protein analysis work. The useful thing about having the whole genome of Homotherium is that you can of course simply translate the sequence data to work out what proteins would be made.
Hopefully more work on Stephanorhinus is planned; it would be really interesting to compare proteins of S. hemitoechus, S. kirchbergensis, S. hundsheimensis and Coelodonta antiquitatis to see their inter-relationships.
Also given what ancient DNA says about hybridisation in the ancestry of Palaeoloxodon it wolud be interesting to sequence proteins of Elephas jolensis; Loxodonta atlantica; and several stages of P/E. reckii and compare them to those of P. antiquus, P. namadicus and P. naumanni as well as Mammuthus and all living elephants (as well as any dwarfed mediterranean elephants available).
And of course where Elasmotherium sibiricum sits amongst all those rhinos too.
oh yes definitely.
How exciting. I can’t tell you how thrilled I was to read about this study as Homotherium has always been a favorite.
Hopefully we’ll get coat colour and appearance genes yet. For now do you are think we can infer a tawny coat based on their probable habitat and hunting technique?
I need to get onto some sort of RSS for these sorts of papers.
Also, is there anything coming up on P. atrox or Miracinonyx?
I’d like to see where the Asiatic Linsang sits on this chart. Between hyaena and the cats, I think.
Yes, that’s where I would expect too. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0234385 https://www.sciencedirect.com/science/article/pii/S0960982217311983
Pingback: The most (and least) read posts of 2021 | TwilightBeasts